A brief summary of the manuscript, Antimicrobial-resistant bacterial populations and antimicrobial resistance genes obtained from environments impacted by livestock and municipal waste (Agga et al., 2015)
Key Points
- Antibiotic resistance is naturally occurring in almost any environment, making it difficult to determine what is the true human health risk associated with any change in environmental bacterial populations or genes.
- This study found higher concentrations of resistant bacteria in both treated municipal wastewater and livestock wastes than are naturally present in soil or water.
- Municipal wastewater samples contained a wider variety of antibiotic-resistant genes than were present in livestock wastes.
Summary
Research has found that antibiotic resistance occurs naturally in bacteria living in almost any environment, including in places with little or no human contact, such as soils under native, ungrazed prairies, or in the Antarctic. Antibiotic resistance has also been found in intestinal tracts of humans and animals that have not been exposed to antibiotics.
However, the increasing number of antibiotic-resistant pathogens is still a serious health concern, and antibiotic residues released to the environment from food animal production, horticulture, and municipal waste (treated sewage) may contribute to the problem. To test the level of antibiotic resistance that can be attributed to human or animal waste, a 2015 study by researchers at the U.S. Department of Agriculture looked at the number of antibiotic-resistant bacteria and the number of DNA segments (genes) that are known to grant resistance to bacteria that were present in the effluent (discharge) from municipal wastewater treatment plants, swine lagoons, beef cattle feedlot runoff ponds, and from two environments expected to have little or no impact from human or animal waste (pristine lake water and prairie soil).
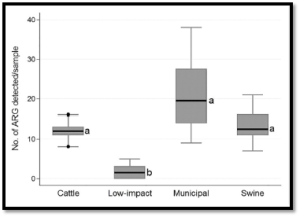
The study found that samples from livestock environments had higher concentrations of resistant live bacteria than were present in treated municipal wastewater, which one would expect since wastewater treatment plants are designed to reduce the number of fecal bacteria in sewage prior to discharge into a river or stream. Also not too surprising was that both the treated effluent and livestock wastes had significantly higher levels of resistant bacteria than were present in the environments that normally had no human or livestock presence [‘Low-impact’]. However, when the researchers looked at the DNA results they found that municipal effluent contained a larger number and wider variety of genes coding for resistance to what would be considered clinically important, meaning resistance to a type of antibiotic that is used in humans. This is not surprising if we consider that fully a third of antibiotics used in animals (ionophores) have no equivalence in human medicine; and resistance to an additional 40% of animal use antibiotics (tetracyclines) were represented as only 2 of the 62 genes tested in this study.
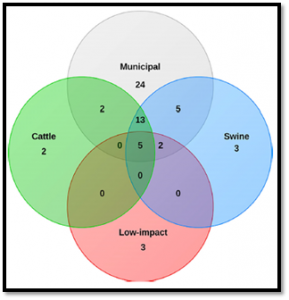
The researchers took care to note that while antibiotic resistance is higher in animal waste and municipal sewage compared to background levels, it is difficult to determine the human health impact of this increase because antibiotic resistance is natural to the environment making tracking very difficult. So, if future research does find that increasing antibiotic resistance in environmental bacteria poses a risk for human health, then changes may be needed in how both municipal and animal wastes are handled in order to reduce their contributions of antibiotic resistance to the environment.
Ready for a deeper dive?
Learn more about the potential impact of antibiotic resistance in human and animal wastes:
Authors
Written by Mara Zelt while a Graduate Student in the Dept. of Biological Systems Engineering, University of Nebraska. Reviewed by: Rick Stowell, University of Nebraska and Jovana Kovacevic, Oregon State University
The scientific research summarized in this article was published as:
Agga, G. E., Arthur, T. M., Durso, L. M., Harhay, D. M., Schmidt, J. W. (2015). Antimicrobial-Resistant Bacterial Populations and Antimicrobial Resistance Genes Obtained from Environments Impacted by Livestock and Municipal Waste. PLOS ONE, 10(7), e0132586. https://doi.org/10.1371/journal.pone.0132586 .
The article presents the author’s interpretation of the published research for a general audience and should not be considered a reflection of the position or opinion of the researchers.

