A summary of The Human Health Implications of Antibiotic Resistance in Environmental Isolates from Two Nebraska Watersheds by Donner et al. 2022
Key points
- The interconnected health of humans, animals, and the environment is well established and increasingly studied in concert within a “One-Health” framework.
- Approximately 40% of the bacteria isolated from watersheds in this study had acquired new antibiotic resistance genes which they had picked up in the environment.
- Both urban and agricultural watersheds contained antibiotic-resistant bacteria, demonstrating the importance of one-health-based decision-making across industries and institutions.
Summary
Bacteria are found everywhere and are mostly harmless. Some are even important for digestion or for producing yogurt and cheese. However, other bacteria can cause infections that can make people sick, and are often treated with antibiotics. These antibiotics do a good job at killing most bacteria, but other bacteria develop protection against them, known as antibiotic resistance. The global use of antibiotics to treat infections is connected to the overall health of humans, animals, and the environment. This concept is known as One-Health. In addition to treating humans, antibiotics are used in livestock production and specialty crops such as orchards. The antibiotics pass through their respective systems and end up in wastewater, soils, and streams where environmental bacteria can also develop resistance. One of the main goals of the One-Health initiative is to decrease the chances of disease-causing bacteria developing resistance to antibiotics that are important in human and animal medicine.

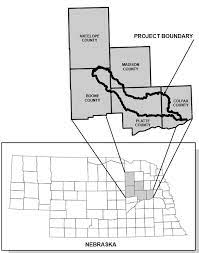
Researchers from the University of Nebraska – Lincoln evaluated the link between antibiotics and antibiotic-resistant bacteria in stream water. The researchers took multiple water samples from points along the waterways in two watersheds. The Elkhorn River watershed drains an area made up of agricultural land and urban areas with contributions from wastewater treatment plants. The other is the Shell Creek watershed, which drains an area dominated by agriculture.
The researchers detected six antibiotics of human importance in the surface water from the two watersheds. This shows that we can detect the use of antibiotics in upstream areas like hospitals or livestock production in downstream rivers. This is an important finding because antibiotics can pressure bacteria in the environment to develop antibiotic resistance.
The researchers grew bacteria from the water samples and were able to isolate bacteria capable of causing human infection. These potentially disease-causing bacteria were further tested to see if they were resistant to specific antibiotics. The researchers measured the DNA of the isolated bacteria and identified the ways that the bacteria protect themselves from being killed by the detected antibiotics. These protections are known as antibiotic resistance genes. The researchers found that around 40% of the bacteria isolated from both watersheds acquired new antibiotic resistance genes from their surroundings. This indicates that antibiotic resistance genes are moving between bacteria that originated from different sources such as humans and animals and end up mixed together in the environment.
This study’s results highlight the importance of a One Health approach to assessing the spread of antibiotic resistance. For example, antibiotics in hospitals contribute to the same pool of antibiotic resistance as does the use of antibiotics in livestock production. Realizing this will lead to the more careful and responsible use of antibiotics as we learn more about how human, animal, and environmental health are all connected.
Ready to learn more?
Learn more about AMR from a One-Health perspective:
- One-health basics
- Advancing the One Health response to Antimicrobial Resistance (AMR)
- Antibiotic Resistance: One Health One World Outlook
Authors
Written by Timothy Neher, graduate research assistant, Iowa State University.
The scientific research summarized in this article was published as:
Donner L, Staley ZR, Petali J, Sangster J, Li X, Mathews W, et al. The Human Health Implications of Antibiotic Resistance in Environmental Isolates from Two Nebraska Watersheds. Microbiol Spectr. 2022; https://doi.org/10.1128/spectrum.02082-21
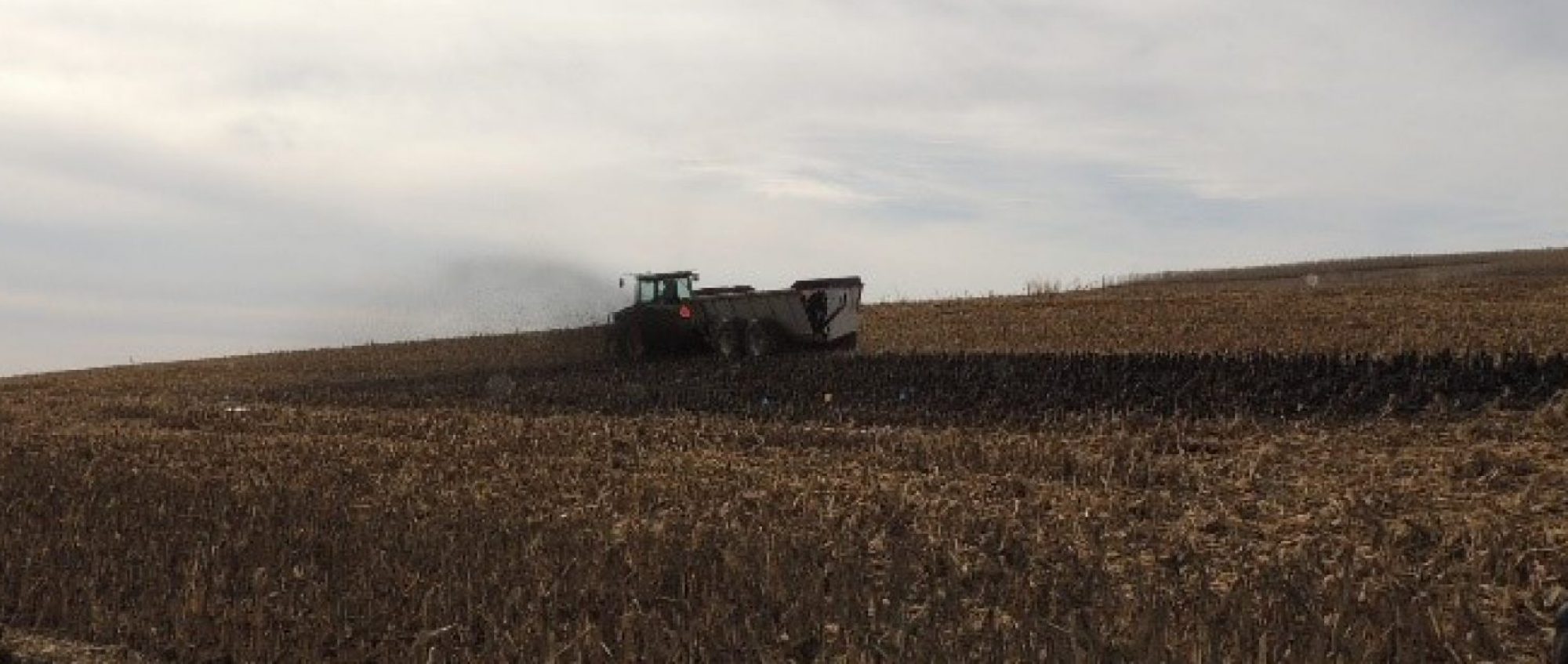
Photo credits: Elkhorn River; Shell Creek

