A summary of Using sewage for surveillance of antimicrobial resistance by Aarestrup and Woolhouse (2020)
Key Points
- Sewage-based surveillance for antimicrobial resistance provides a flexible, scalable, and quickly implementable AMR tracking method.
- Advances in DNA sequencing enable faster and more responsive resistance monitoring, which is essential to address AMR surveillance worldwide.
Summary
The world is learning more about the increasing threat that antimicrobial resistance (AMR) has on global health. More people are being hospitalized, and dying because of microbial (bacteria, fungi, viral, etc.) infections that are now resistant to antimicrobial treatments that were effective in the past. AMR is a serious and systemic problem that will require broadly shared action on multiple fronts to address the crisis. One essential part of combating AMR is developing a reliable form of surveillance to track AMR progression over time. This could be effectively done through surveillance of sewage. After all, everyone must ‘go’ sometimes, it is only natural, and when we go our feces contains millions of microbes that offer a valuable window into the microbial life in our bodies, including the level of antimicrobial resistance.
The mass of microbial life living in human waste is more than a window into our health, it also has historically been a pathway for passing disease and that is why the advent of sewage treatment has been one of the most important inventions for human health in world history; and with a growing world population increasingly connected to sewage systems, sewage surveillance for tracking community health has many advantages. Testing sewage is a cheaper, more efficient, and non-invasive way to sample large urban populations. Some benefits to surveying sewage are that implementation is straightforward and requires less expensive equipment for sample collection. Fewer samples are needed to provide representative results, compared to clinical surveillance for AMR. Moreover, the methods for sewage surveillance can be more easily standardized, and the results have been shown to mirror AMR in the community discovered through traditional clinical surveillance. Sewage is already recommended to screen for polio, so adding AMR surveillance is a reasonable extension to assess the general level of resistance in a community. The one potential drawback, that the tests cannot be linked back to any individual, does have the benefit of providing anonymous results, which reduces the logistical or regulatory burden that is often associated with tracing in human health data.
The idea of broadly testing for AMR is not new, the World Health Organization founded their Global Antimicrobial Resistance Surveillance System (GLASS) in 2015 to foster national surveillance of AMR and track emerging resistance and global spread. However, most of the samples collected in these programs are from hospitalized patients on last-resort antimicrobial treatments. This means that the sample sizes are small and not representative of the level of resistance that would be expected in the general population. Additionally, the surveillance methods used in these programs in the past were not easily done outside of a medical laboratory and required expensive equipment not typically available at a municipal wastewater treatment plant. Thus, there has been some difficulty combining results for different kinds of tests. These issues have made tracing AMR in the past more challenging.
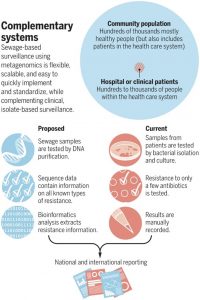
Sewage surveillance could potentially solve some of the issues previously encountered while tracking AMR and utilize advances in genomic sequencing which allows for cheaper and faster sequencing and determination of AMR genes in the sewage. There are some necessary steps to put sewage surveillance into action broadly. Protocols will need to be developed to make sure the data collected is standardized, allowing different test locations across the globe to compare their findings. These protocols should be open to modification as new methods and technologies become available. Sewage surveillance alone is not enough to manage the oncoming antimicrobial resistance crisis and it should augment current tracking and surveillance systems. Sewage surveillance is underutilized relative to the cost of its implementation; including it will add to the population-based information on AMR surveillance from federal and local health agencies in the US and worldwide.
Ready to learn more?
Learn more about AMR in wastewater systems:
- Water and sanitation: an essential battlefront in the war on antimicrobial resistance
- Antibiotics and resistance genes in wastewater treatment plants
- WHO briefing note on emerging water, sanitation, and hygiene issues related to AMR
Authors
Written by Andrew Lutt while an Undergraduate Researcher in the Dept. of Biological Systems Engineering, University of Nebraska. Reviewed by: Noelle Mware, University of Nebraska, and Carlton Poindexter, University of Maryland.
The scientific research summarized in this article was published as:
Aarestrup, F.M and Woolhouse, M.E.J. (2020). Using sewage for surveillance of antimicrobial resistance. Science, 367(6478). DOI: 10.1126/science.aba3432
The article presents the author’s interpretation of the published research for a general audience and should not be considered a reflection of the position or opinion of the researchers.

